Research Summary
Electron Tunneling in Respiratory Complex I:
Alexei Stuchebrukhov Group, UC, Davis (2008-present)
Developed a code to calculate the electron tunneling pathways at atomic
resolution, tunneling matrix elements, and electron transfer rates of
long-distance electron tunneling in proteins. The code works in
conjunction with Gaussian quantum chemistry package,
and can thus be applicable at any computational level including density functional theory.
The system studied is complex I (NADH:ubiquinone
oxidoreductase), a key enzyme in the
respiratory
electron transport chain.
Complex I couples the transfer of electrons over 90 Å along seven Fe/S clusters from NADH to ubiquinone to the creation of the proton gradient across the membrane necessary for ATP synthesis.
Complex I defects are the cause of many
neurodegenerative diseases including Parkinson's disease, Alzheimer's disease, and
Huntington's disease. We for the
first time identified distinct tunneling pathways, key
protein residues, and proposed a unique role of internal water as a
mediator [1]
Realistic Simulation of Anharmonic Vibrational Dynamics in Solution:
Shaul Mukamel Group, UC, Irvine (2003-2008)
Developed a new methodology for computationally efficient realistic
simulation of the anharmonic vibrational dynamics of polyatomic
molecules in solution (as an extension of my work in the graduate
course in
Hiro-o Hamaguchi Group [2]). The
model can treats vibrational dynamics such as dephasing, population
transfer, exciton hopping, and correlated frequency fluctuations of
multiple-quanta transitions of molecules as large as proteins based on
a QM/MM scheme without repeating quantum mechanical (QM) calculations
at each time step. The ab initio anhrmonic multi-dimensional
vibrational Hamiltonian of QM regions (vibrational chromophores) is
parameterized with respect to the local multipole electric field
(denoted as Electronic DFT Maps) [3]. Coulombic interactions
between multiple QM
regions are expanded in transition multipoles [4].
By combining them with MD trajectories of the whole system
and
wave-propagation techniques, the vibrational dynamics of the whole
system is obtained. The proposed technique is one of the most accurate
and has been successfully applied to water (HOD/D2O and pure H2O) and
many proteins, in good agreement with experiment. Simulations of the
whole four amide vibrational regions (I, II, III and A) of proteins
were performed for the first time [3].
Coherent 2-Dimenstional Infrared Spectroscopy (2DIR) of Biomolecules
Shaul Mukamel Group, UC, Irvine (2003-2008)
Applied the proposed technique to coherent 2-Dimensinal infrared (2DIR)
spectrsocopy. 2DIR has been established as a powerful tool to
investigate the molecular structure and ultrafast molecular events
due to its unique high
temporal resolution and multidimensional spectral information, although
spectral analysis had been difficult because of the complex nature of
the
vibrational Hamiltonian.
We identified the 2DIR signatures of the extraordinary
high
anharmonicity of the malonaldehyde (a double-well type intramolecular
proteon transfer system) [5], correlations and
excitonic
couplings in the
hydrogen-bond network of liquid water, correlated hydrogen bond
formation at peptide amide bonds [6], and protein secondary
structures.
We demonstrated that 2DIR with polarized beams can distinguish multiple
secondary structure candidates of a peptide which NMR cannot
distinguish.[7]
- Tomoyuki Hayashi, and
Alexei Stuchebrukhov, Proc.
Natl. Acad. Sci. USA, 107,
19157 (2010) [pdf]
- Tomoyuki Hayashi, and
Hiro-o Hamaguchi, Chem.
Phys. Lett. 326, 115 (1998) [pdf]
- Tomoyuki Hayashi, Wei
Zhuang and Shaul Mukamel, J. Phys. Chem. A109,
9747 (2005) [pdf]
- Tomoyuki Hayashi, and
Shaul Mukamel, J. Phys. Chem. B 111,
11032 (2007) [pdf]
- Tomoyuki Hayashi and
Shaul Mukamel, J. Phys. Chem. A 107, 9113 (2003) [pdf]
- Tomoyuki Hayashi, and
Shaul Mukamel, J. Chem. Phys.
125, 194510 (2006)
[pdf]
- Wei Zhuang, Tomoyuki Hayashi,and Shaul Mukamel,
Angew. Chem.
48, 3750 (2009) [pdf]
Search
News
- (2010/11) Our study on "Electron Tunneling in Respiratory Complex I" is published in PNAS.
[pdf]
Contact Info
Tomoyuki Hayashi
Chemistry Department
University of California, Davis
One Shields Ave
Davis, CA 95616, USA
e-mail: thayashi@ucdavis.edu
Copyright © 2010-2011 Tomoyuki Hayashi | Last Modified 25 Jan 2011
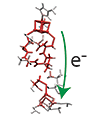
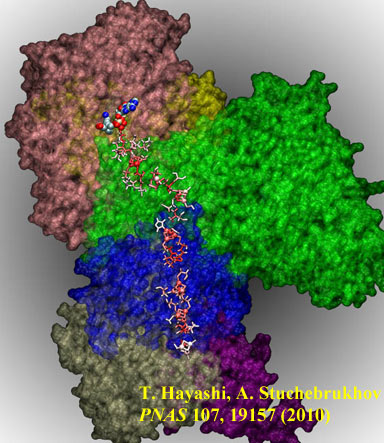
Developed a code to calculate the electron tunneling pathways at atomic resolution, tunneling matrix elements, and electron transfer rates of long-distance electron tunneling in proteins. The code works in conjunction with Gaussian quantum chemistry package, and can thus be applicable at any computational level including density functional theory.
The system studied is complex I (NADH:ubiquinone oxidoreductase), a key enzyme in the respiratory electron transport chain. Complex I couples the transfer of electrons over 90 Å along seven Fe/S clusters from NADH to ubiquinone to the creation of the proton gradient across the membrane necessary for ATP synthesis. Complex I defects are the cause of many neurodegenerative diseases including Parkinson's disease, Alzheimer's disease, and Huntington's disease. We for the first time identified distinct tunneling pathways, key protein residues, and proposed a unique role of internal water as a mediator [1]
Realistic Simulation of Anharmonic Vibrational Dynamics in Solution:
Shaul Mukamel Group, UC, Irvine (2003-2008)
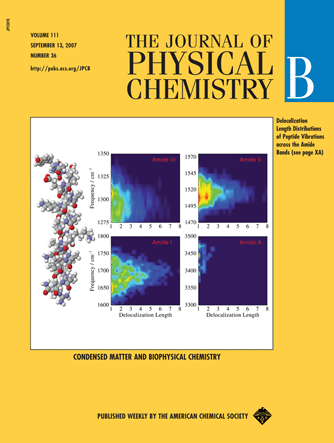
Developed a new methodology for computationally efficient realistic simulation of the anharmonic vibrational dynamics of polyatomic molecules in solution (as an extension of my work in the graduate course in Hiro-o Hamaguchi Group [2]). The model can treats vibrational dynamics such as dephasing, population transfer, exciton hopping, and correlated frequency fluctuations of multiple-quanta transitions of molecules as large as proteins based on a QM/MM scheme without repeating quantum mechanical (QM) calculations at each time step. The ab initio anhrmonic multi-dimensional vibrational Hamiltonian of QM regions (vibrational chromophores) is parameterized with respect to the local multipole electric field (denoted as Electronic DFT Maps) [3]. Coulombic interactions between multiple QM regions are expanded in transition multipoles [4].
By combining them with MD trajectories of the whole system and wave-propagation techniques, the vibrational dynamics of the whole system is obtained. The proposed technique is one of the most accurate and has been successfully applied to water (HOD/D2O and pure H2O) and many proteins, in good agreement with experiment. Simulations of the whole four amide vibrational regions (I, II, III and A) of proteins were performed for the first time [3].
Coherent 2-Dimenstional Infrared Spectroscopy (2DIR) of Biomolecules
Shaul Mukamel Group, UC, Irvine (2003-2008)
Applied the proposed technique to coherent 2-Dimensinal infrared (2DIR)
spectrsocopy. 2DIR has been established as a powerful tool to
investigate the molecular structure and ultrafast molecular events
due to its unique high
temporal resolution and multidimensional spectral information, although
spectral analysis had been difficult because of the complex nature of
the
vibrational Hamiltonian.
We identified the 2DIR signatures of the extraordinary
high
anharmonicity of the malonaldehyde (a double-well type intramolecular
proteon transfer system) [5], correlations and
excitonic
couplings in the
hydrogen-bond network of liquid water, correlated hydrogen bond
formation at peptide amide bonds [6], and protein secondary
structures.
We demonstrated that 2DIR with polarized beams can distinguish multiple
secondary structure candidates of a peptide which NMR cannot
distinguish.[7]
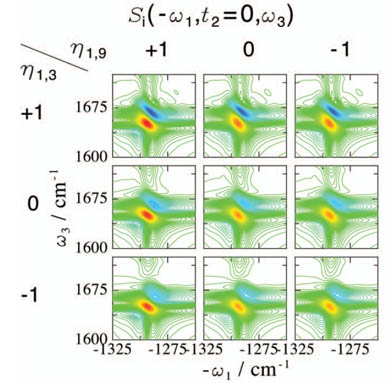
Applied the proposed technique to coherent 2-Dimensinal infrared (2DIR) spectrsocopy. 2DIR has been established as a powerful tool to investigate the molecular structure and ultrafast molecular events due to its unique high temporal resolution and multidimensional spectral information, although spectral analysis had been difficult because of the complex nature of the vibrational Hamiltonian.
We identified the 2DIR signatures of the extraordinary high anharmonicity of the malonaldehyde (a double-well type intramolecular proteon transfer system) [5], correlations and excitonic couplings in the hydrogen-bond network of liquid water, correlated hydrogen bond formation at peptide amide bonds [6], and protein secondary structures. We demonstrated that 2DIR with polarized beams can distinguish multiple secondary structure candidates of a peptide which NMR cannot distinguish.[7]
- Tomoyuki Hayashi, and Alexei Stuchebrukhov, Proc. Natl. Acad. Sci. USA, 107, 19157 (2010) [pdf]
- Tomoyuki Hayashi, and Hiro-o Hamaguchi, Chem. Phys. Lett. 326, 115 (1998) [pdf]
- Tomoyuki Hayashi, Wei Zhuang and Shaul Mukamel, J. Phys. Chem. A109, 9747 (2005) [pdf]
- Tomoyuki Hayashi, and Shaul Mukamel, J. Phys. Chem. B 111, 11032 (2007) [pdf]
- Tomoyuki Hayashi and Shaul Mukamel, J. Phys. Chem. A 107, 9113 (2003) [pdf]
- Tomoyuki Hayashi, and Shaul Mukamel, J. Chem. Phys. 125, 194510 (2006) [pdf]
- Wei Zhuang, Tomoyuki Hayashi,and Shaul Mukamel,
Angew. Chem.
48, 3750 (2009) [pdf]
Search
News
- (2010/11) Our study on "Electron Tunneling in Respiratory Complex I" is published in PNAS. [pdf]
Contact Info
Tomoyuki HayashiChemistry Department
University of California, Davis
One Shields Ave
Davis, CA 95616, USA e-mail: thayashi@ucdavis.edu